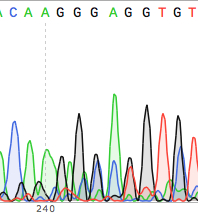
This post is essentially a plea to the internet for someone to tell me there's a better program/method out there!. This combination is fairly straightforward when the sequencing confirms everything nicely, but when there's been a problem somewhere it takes a lot of time and effort to highlight it. 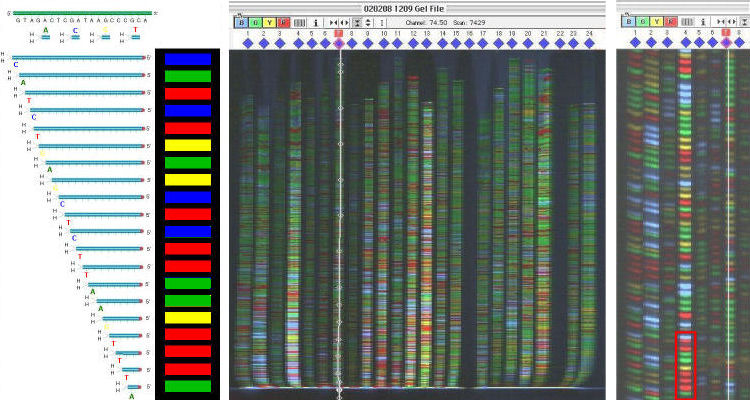
Reverse Complement ( ) - for RCing sequencesĬlustal Omega ( ) - to align my sequencing to the sequences I have in my maps and check it's correct or notĮxPASy ( ) - to check the sequenced data is 'in frame' and that the translated protein sequence is how it should beīLAST ( ) - last resort in bizarre cases of "wtf is this random sequence in the middle of things?" SeqTrace is a new, free, and open-source software application that is designed to automate the entire workflow by facilitating easy batch processing of large numbers of trace files. Microsoft Word - to handle raw/FASTA sequences Snapgene (free) viewer - for plasmid map creation/visualisation/restriction site mapping/chromatogram viewer My question is: What programs do people recommend for looking at your sequencing data? It's my first time experiencing cloning without being babysat really and the struggle appears with the sequencing data. Creating a few simple constructs of fluorescent reporter tagged proteins. There is also no way to view the raw data signal.I'm a first year PhD Biochemistry student and I've been doing some cloning recently, fairly simple stuff, just ligating sequences into a 3.1 hygro vector. A major downside of 4Peaks is that the quality score bars are shown in the background which creates confusion when viewing the quality and chromatogram in the same window. Traces appear sharper when viewed through 4Peaks compared to other chromatogram viewing programs, which allows for a easy viewing. CostĤPeaks boasts that it is able to “render peaks better than anyone else”. One handy feature is the speech function where 4Peaks will verbally read out the sequence of the file so you can follow along while looking at another sequence to compare differences. You also have the ability to translate the DNA sequence into the amino acid sequence across 3 separate frames. The program is full of nifty features, you have the ability to open a separate txt window containing the sequence and select from the peaks/txt file at the same time to compare specific regions. This function is not available in 4Peaks. ABI limits (regions outside of clear range region are displayed in grey) It is possible to edit, insert and delete bases in 4Peaks. 4 peaks for Macintosh OSX is available at: Options for PC are available at: Chromas is avalable at: Sequence Scanner Software v1. However, it will take you into a separate web browser. You can run a BLAST search through 4Peaks. Click on the appropriate icon (s) to go to the respective Web page. It is possible to manually select areas to trim at the 3′ and 5′ ends. A number of free software programs are available for viewing trace or chromatogram files. Through the quality overview function you can trim below a set quality value. Input file formatsĪBI, SCF, ALF, PLN, EXP, ZTR, BIO and TXT. Modern applications of Sanger DNA sequencing often require converting a large number of chromatogram trace files into high-quality DNA sequences for. 4Peaks does not contain any features for DNA alignment or assembly. There is now way to view the raw data in 4Peaks. Quality scores are presented as blue bars behind the peaks and at the top of the window when a peak is selected. Supported platforms: MacOSX 10.7 or later DNA Sequencing Reaction Clean-up using Phenol & Butanol.Tris EDTA DNA Sequencing Resuspension Buffer.Exonuclease I – Shrimp Alkaline Phosphatase Clean Up of PCR Products.Auto PeakTrace 6 Online Activation Guide.How to Update the PeakTrace License on Linux.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
